2 Perspectives on the phylogenetic tree
Phylogenetic modelling concepts are constantly changing. It is one of the most dynamic fields of study in all biology. Over the last several decades, new research has challenged scientists’ ideas about how organisms are related. The scientific community has proposed new models of these relationships.
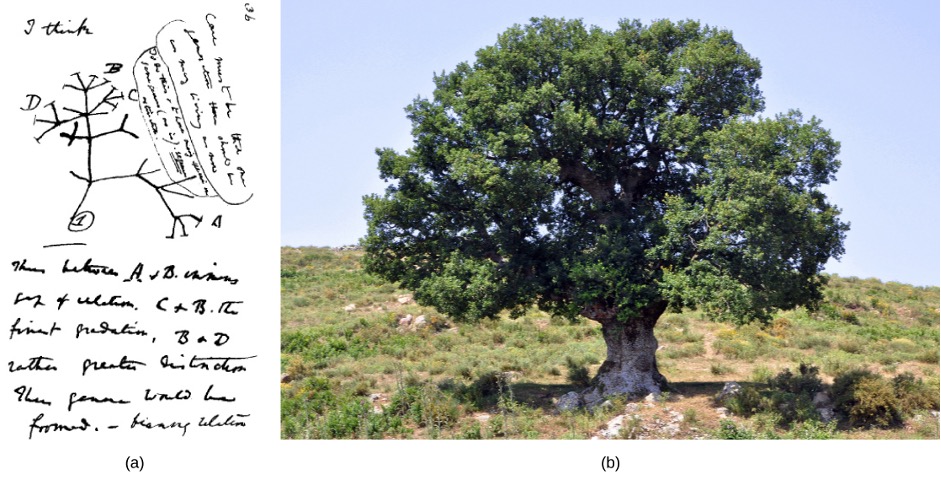
Many phylogenetic trees are models of the evolutionary relationship among species. Phylogenetic trees originated with Charles Darwin, who sketched the first phylogenetic tree in 1837 (Figure 1.8a). This served as a prototype for subsequent studies for more than a century. The phylogenetic tree concept with a single trunk representing a shared ancestry, with the branches representing the divergence of species from this ancestry, fits well with the structure of many common trees, such as the oak (Figure 1.8b). However, evidence from modern DNA sequence analysis and newly developed computer algorithms has caused scepticism about the standard tree model’s validity in the scientific community.

Classical thinking about prokaryotic evolution, included in the classic tree model, is that species evolve clonally. That is, they produce offspring themselves with only random mutations causing the descent into the variety of modern-day and extinct species known to science. This view is somewhat complicated in eukaryotes that reproduce sexually, but the laws of Mendelian genetics explain the variation in offspring, again, to be a result of a mutation within the species. Scientists did not consider the concept of genes transferring between unrelated species as a possibility until relatively recently. Horizontal gene transfer (HGT), or lateral gene transfer, is the transfer of genes between unrelated species. HGT is an ever-present phenomenon, with many evolutionists postulating a major role for this process in evolution, thus complicating the simple tree model. Genes pass between species which are only distantly related using standard phylogeny, thus adding a layer of complexity to understanding phylogenetic relationships.
Genome fusion and eukaryote evolution
Scientists believe the ultimate in HGT occurs through genome fusion between different prokaryote species when two symbiotic organisms become endosymbiotic. This occurs when one species is taken inside another species’ cytoplasm, which ultimately results in a genome consisting of genes from both the endosymbiont and the host. This mechanism is an aspect of the Endosymbiont Theory, which most biologists accept as the mechanism whereby eukaryotic cells obtained their mitochondria and chloroplasts. However, the role of endosymbiosis in developing the nucleus is more controversial. Scientists believe that nuclear and mitochondrial DNA have different (separate) evolutionary origins, with the mitochondrial DNA being derived from the bacteria’s circular genomes engulfed by ancient prokaryotic cells. We can regard mitochondrial DNA as the smallest chromosome. Interestingly enough, mitochondrial DNA is inherited only from females. The mitochondrial DNA degrades in sperm when the sperm degrades in the fertilised egg or in other instances when the mitochondria located in the sperm’s flagellum fails to enter the egg.
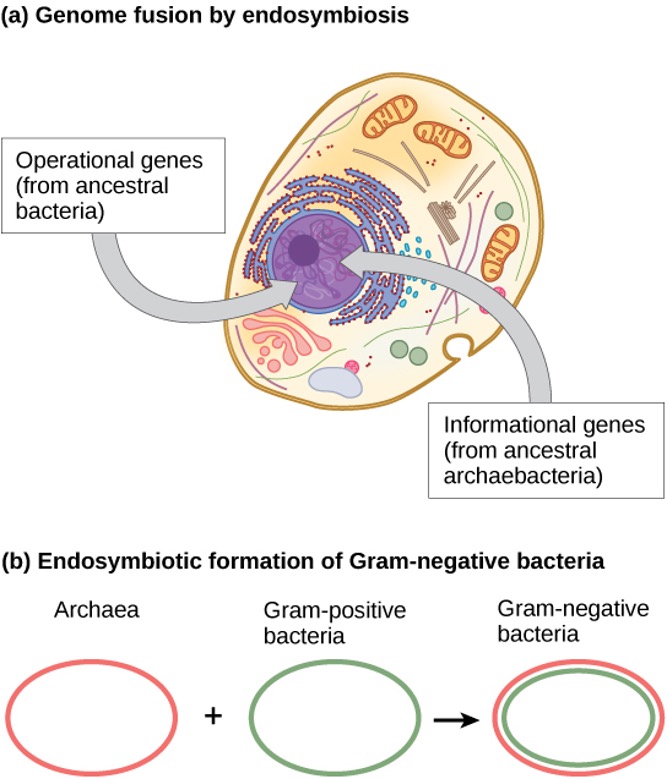
Within the past decade, James Lake of the UCLA/NASA Astrobiology Institute proposed that the genome fusion process is responsible for the evolution of the first eukaryotic cells (Figure 1.9a). Using DNA analysis and a new mathematical algorithm, conditioned reconstruction (CR), his laboratory proposed that eukaryotic cells developed from an endosymbiotic gene fusion between two species, one an Archaea and the other a Bacteria. As mentioned, some eukaryotic genes resemble those of Archaea; whereas, others resemble those from Bacteria. An endosymbiotic fusion event, such as Lake has proposed, would clearly explain this observation. Alternatively, this work is new and the CR algorithm is relatively unsubstantiated, which causes many scientists to resist this hypothesis.
Lake’s more recent work (Figure 1.9b) proposes that gram-negative bacteria, which are unique within their domain in that they contain two lipid bilayer membranes, resulted from an endosymbiotic fusion of archaeal and bacterial species. The double membrane would be a direct result of the endosymbiosis, with the endosymbiont picking up the second membrane from the host as it was internalised. Scientists have also used this mechanism to explain the double membranes in mitochondria and chloroplasts. Some are sceptical of Lake’s work, and the biological science community still debates his ideas. In addition to Lake’s hypothesis, there are several other competing theories as to the origin of eukaryotes. How did the eukaryotic nucleus evolve? One theory is that the prokaryotic cells produced an additional membrane that surrounded the bacterial chromosome. Some bacteria have the DNA enclosed by two membranes; however, there is no evidence of a nucleolus or nuclear pores. Other proteobacteria also have membrane-bound chromosomes. If the eukaryotic nucleus evolved this way, we would expect one of the two types of prokaryotes to be more closely related to eukaryotes.
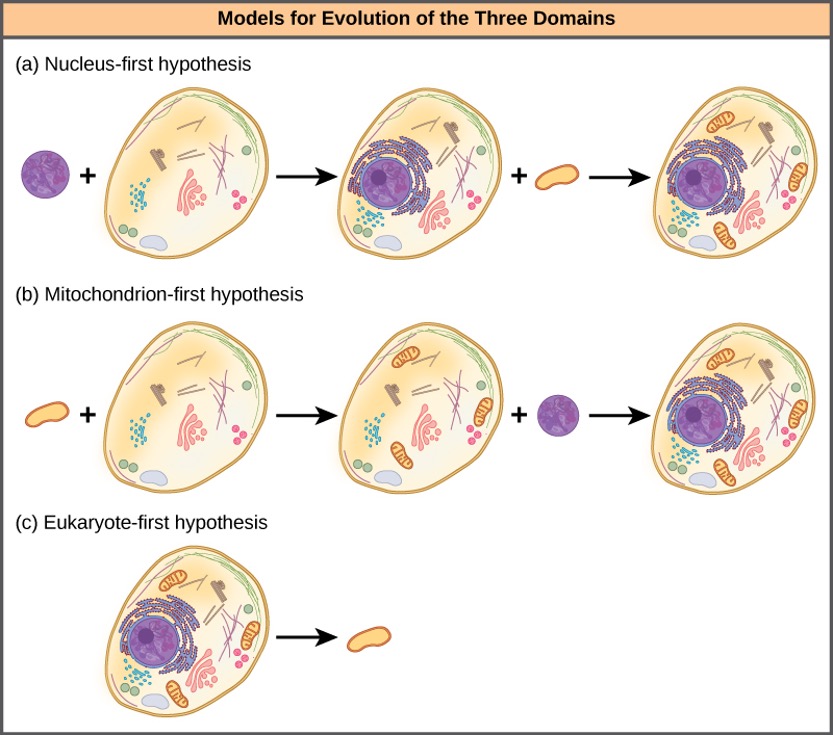
The nucleus-first hypothesis proposes that the nucleus evolved in prokaryotes first (Figure 1.10a), followed by a later fusion of the new eukaryote with bacteria that became mitochondria. The mitochondria-first hypothesis proposes that mitochondria were first established in a prokaryotic host (Figure 1.10b), which subsequently acquired a nucleus, by fusion or other mechanisms, to become the first eukaryotic cell. Most interestingly, the eukaryote-first hypothesis proposes that prokaryotes actually evolved from eukaryotes by losing genes and complexity (Figure 1.10c). All of these hypotheses are testable. Only time and more experimentation will determine which hypothesis data best support.
Web and network models
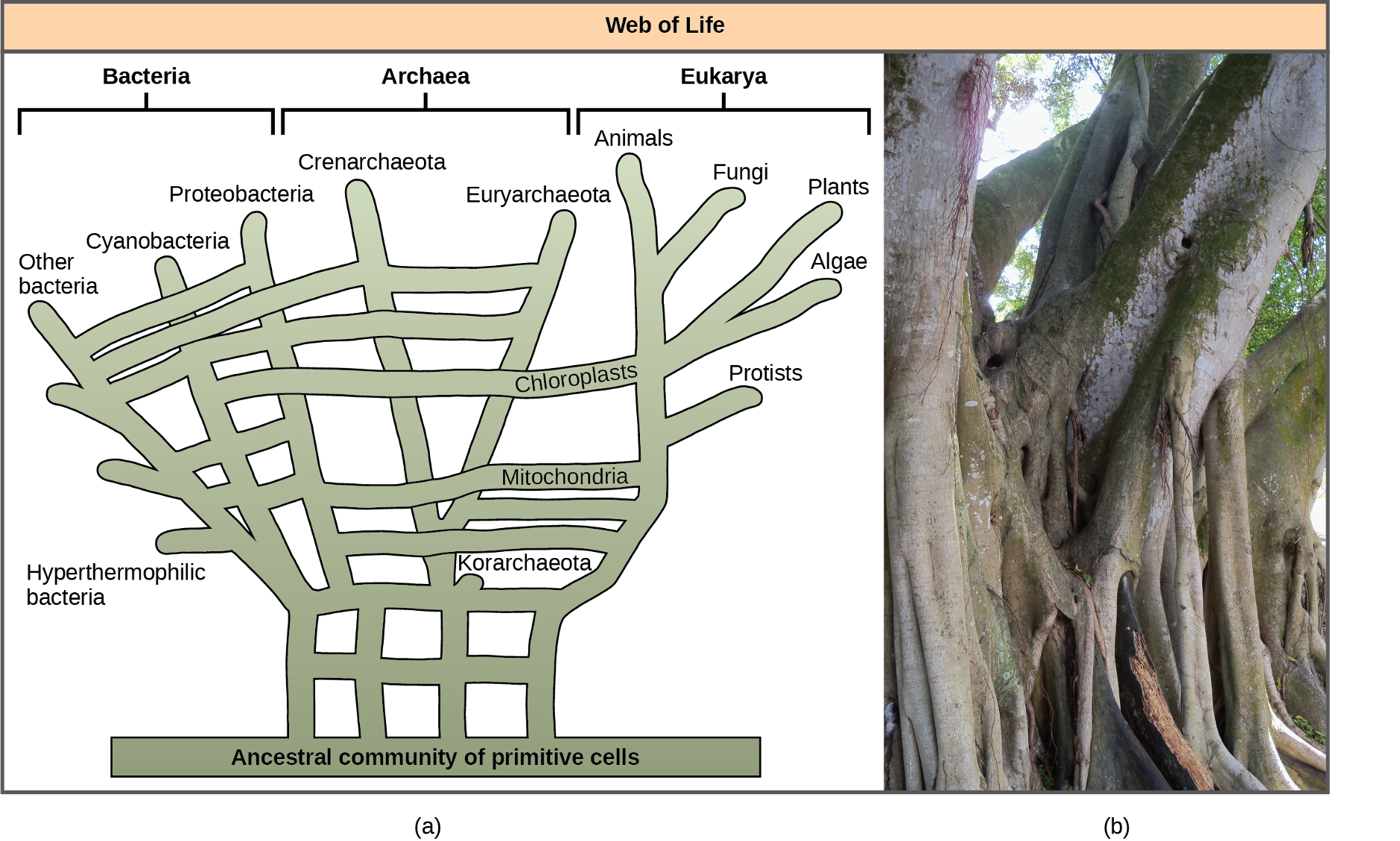
Recognising the importance of HGT, especially in prokaryote evolution, has caused some to propose abandoning the classic “tree of life” model. In 1999, W. Ford Doolittle proposed a phylogenetic model that resembles a web or a network more than a tree. The hypothesis is that eukaryotes evolved not from a single prokaryotic ancestor, but from a pool of many species that were sharing genes by HGT mechanisms. As Figure 1.11a shows, some individual prokaryotes were responsible for transferring the bacteria that caused mitochondrial development to the new eukaryotes; whereas, other species transferred the bacteria that gave rise to chloroplasts. Scientists often call this model the “web of life.” In an effort to save the tree analogy, some have proposed using the Ficus tree (Figure 1.11b) with its multiple trunks as a phylogenetic way to represent a diminished evolutionary role for HGT.
Ring of life models
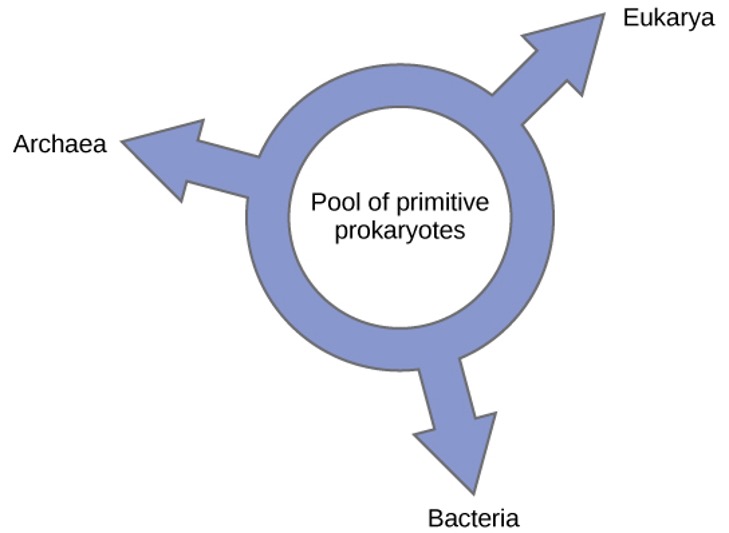
Others have proposed abandoning any tree-like model of phylogeny in favour of a ring structure, the so-called “ring of life” (Figure 1.12). This is a phylogenetic model where all three domains of life evolved from a pool of primitive prokaryotes. Lake, again using the conditioned reconstruction algorithm, proposes a ring-like model in which species of all three domains—Archaea, Bacteria, and Eukarya—evolved from a single pool of gene-swapping prokaryotes. His laboratory proposes that this structure is the best fit for data from extensive DNA analyses performed in his laboratory, and that the ring model is the only one that adequately takes HGT and genomic fusion into account. However, other phylogeneticists remain highly skeptical of this model.
This does not mean a tree, web, or a ring will correlate completely to an accurate description of phylogenetic relationships of life. A consequence of the new thinking about phylogenetic models is the idea that Darwin’s original phylogenetic tree concept is too simple, but made sense based on what scientists knew at the time. However, the search for a more useful model moves on: each model serves as hypotheses to test with the possibility of developing new models. This is how science advances. Researchers use these models as visualisations to help construct hypothetical evolutionary relationships and understand the massive amount of data that requires analysis.

